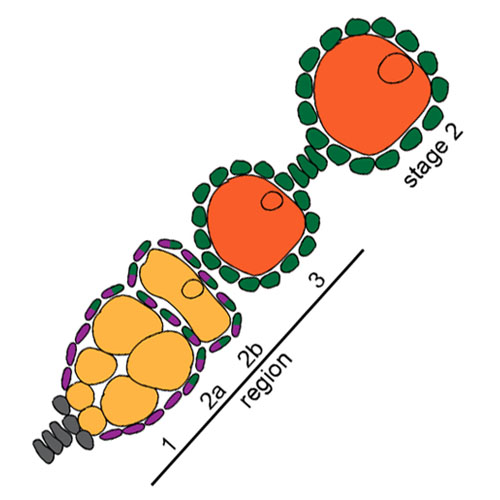
A role for tuned levels of nucleosome remodeler subunit ACF1 during Drosophila oogenesis
02-Feb-2016
Developmental Biology, Volume 411, Issue 2, Pages 217–230, doi:10.1016/j.ydbio.2016.01.039
The Chromatin Accessibility Complex (CHRAC) consists of the ATPase ISWI, the large ACF1 subunit and a pair of small histone-like proteins, CHRAC-14/16. CHRAC is a prototypical nucleosome sliding factor that mobilizes nucleosomes to improve the regularity and integrity of the chromatin fiber. This may facilitate the formation of repressive chromatin. Expression of the signature subunit ACF1 is restricted during embryonic development, but remains high in primordial germ cells. Therefore, we explored roles for ACF1 during Drosophila oogenesis. ACF1 is expressed in somatic and germline cells, with notable enrichment in germline stem cells and oocytes. The asymmetrical localization of ACF1 to these cells depends on the transport of the Acf1 mRNA by the Bicaudal-D/Egalitarian complex. Loss of ACF1 function in the novel Acf17 allele leads to defective egg chambers and their elimination through apoptosis. In addition, we find a variety of unusual 16-cell cyst packaging phenotypes in the previously known Acf11 allele, with a striking prevalence of egg chambers with two functional oocytes at opposite poles. Surprisingly, we found that the Acf11 deletion – despite disruption of the Acf1 reading frame – expresses low levels of a PHD-bromodomain module from the C-terminus of ACF1 that becomes enriched in oocytes. Expression of this module from the Acf1 genomic locus leads to packaging defects in the absence of functional ACF1, suggesting competitive interactions with unknown target molecules. Remarkably, a two-fold overexpression of CHRAC (ACF1 and CHRAC-16) leads to increased apoptosis and packaging defects. Evidently, finely tuned CHRAC levels are required for proper oogenesis.